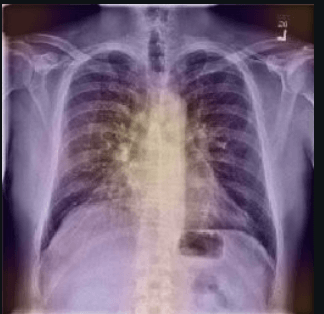
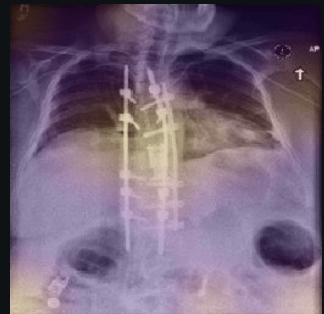
Computer vision has been suggested to help with the battle against COVID-19. In this article, we want to share our preliminary research into two different datasets. Confirming existing results, we found defects with the existing approaches, due to leakage when building COVID image datasets from heterogeneous sources. We explain here why these issues occurred, using interpretability tools to detect them. There is considerable promise for using computer vision to quickly identify COVID-19 and our hope is this will push the development of better image datasets for identifying COVID-19.
According to RSNA, CT Scans are particularly useful in the diagnosis of COVID-19. However, since CT scans are expensive and not readily available everywhere, computer vision experts are looking towards the cheaper and easily accessible x-rays to diagnose COVID-19.
Over the last few weeks, a wide community from researchers to bloggers have suggested using computer vision to identify individuals with COVID-19. They are using the latest tools in AI for computer vision with Convolutional Neural Networks (CNN). The most common idea or solution being proposed is to build a CNN based binary classifier that can classify x-ray images into COVID-19 or Non-COVID-19. CNNs have revolutionized computer vision over the last five years. However, like any AI model, there are two issues that should be considered when using CNN.
- Overfitting or Leakage Exploitation: CNNs can identify completely irrelevant but strong target correlated artifacts in images to predict the Target similar to data leakage in structured datasets.
- Blackbox or Interpretation Limitation: CNNs suffer from interpretability limitations because they rely on complicated nonlinear, multidimensional decision boundaries.
As there is no readily available, well-collated dataset with proper distribution of the target, researchers are cobbling up the training datasets from the undermentioned list of x-ray datasets. They do this by taking the positive class samples from the COVID-19 crowdsourced dataset and the negative class samples from the RSNA and/or Pediatric pneumonia dataset. One of the most known examples of this comes from Researchers from the University of Waterloo and DarwinAI, who have created the COVIDx dataset which builds the dataset by taking positive samples from the COVID-19 crowdsourced dataset and building the negative classes from the Pediatric pneumonia dataset. Based on this, they have built the COVID-Net model to identify COVID-19 positive cases from x-ray images. They have received considerable attention, but also concerns with selection bias.
COVIDx Dataset Analysis
Using DataRobot’s Visual AI, we started by using the COVIDx dataset. We experimented on both Full COVIDx and a downsampled dataset. In the downsampled dataset, we have put together a training dataset using 66 positive COVID results and 120 random selected not COVID examples from the COVIDx dataset. The script to create the dataset is available here.
We built a classifier using a ResNet50 Featurizer on this dataset. The classifier could accurately predict which images were COVID-19 positive. To understand the accuracy, we used interpretability tools such as Image Embeddings and Image Activation Maps. Image Embeddings is an unsupervised approach that shows which images are similar to others. Immediately here, we see two distinct groups, which is a red flag. This means it is trivial to separate these into two classes, COVID-19 positive and COVID-19 negative. The next step is understanding why it is so easy for the classifier to separate these two groups.
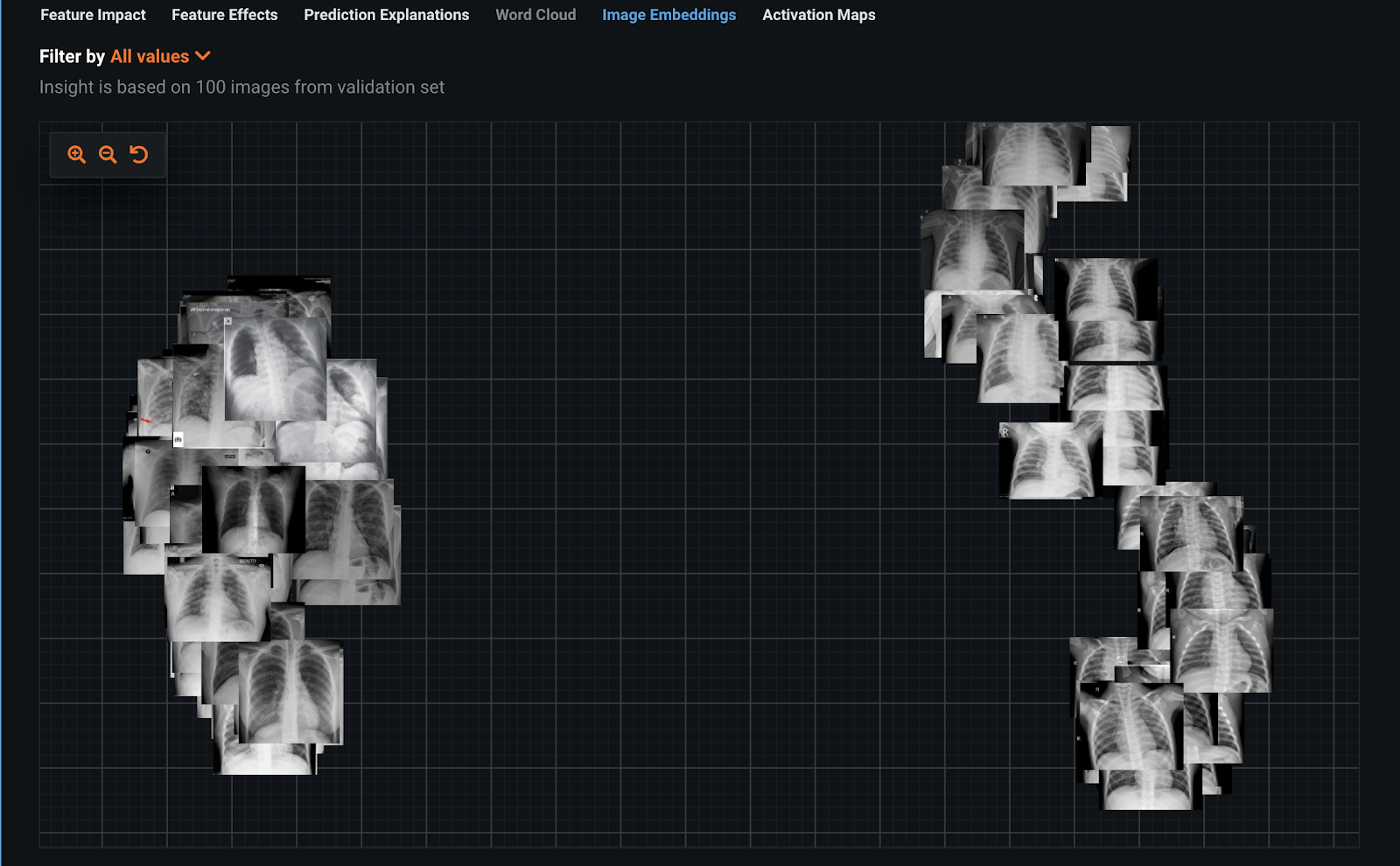
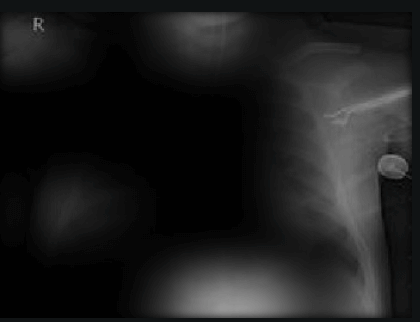
To understand what the image classifier is doing, we can use the interpretability of Activation Maps. Activation Maps show what the model is paying attention to when making its prediction using brightness. Studying these we can identify some examples of leakage.
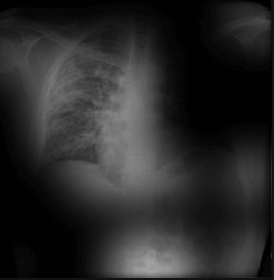
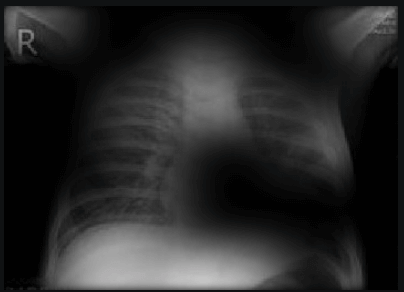
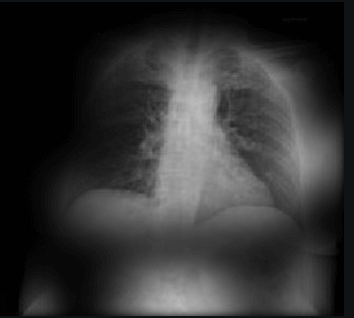
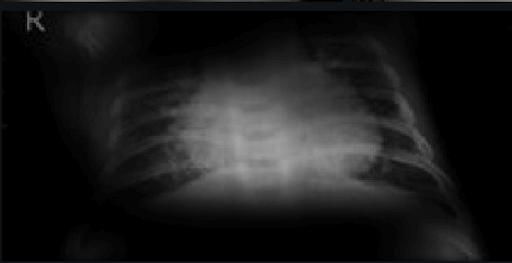
Here the classifier is classifying x-rays with “R” markers by machine as COVID-19 negative as a majority of the x-rays in the COVID-19 negative dataset have these markers. This is a classic example of label leakage, where the AI learns whether or not there is a label. After all, in this case, only images that are COVID-19 negative dataset have a “R” marker.
| COVID-19 Negative |
COVID-19 Positive |
 |
 |
 |
 |
 |
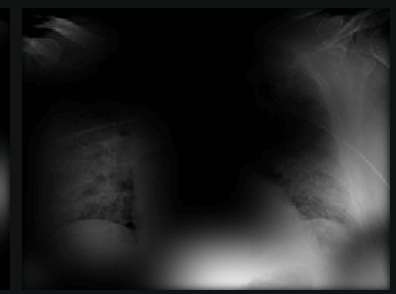 |
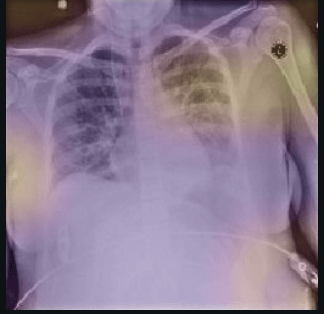
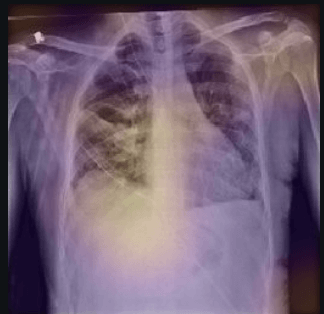
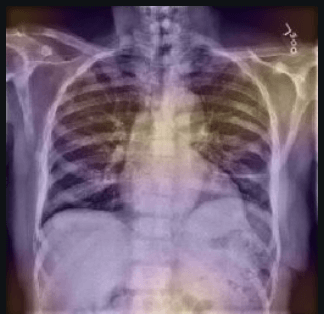
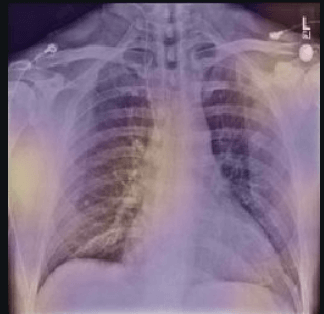
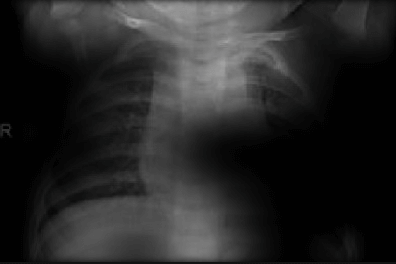
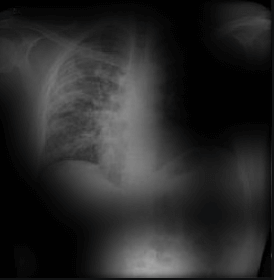
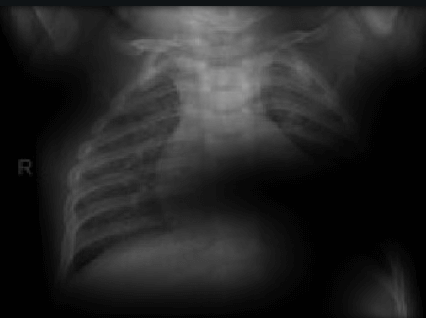
Here the classifier is classifying x-rays with neck, lower jaw, upright shoulder bones and prominent abdominal cavity as COVID-19 negative as a majority of the x-rays in the non-COVID-19 dataset have x-rays with neck, lower jaw, upright shoulder bones and a prominent abdominal cavity.
| COVID-19 Negative |
COVID-19 Positive |
 |
 |
 |
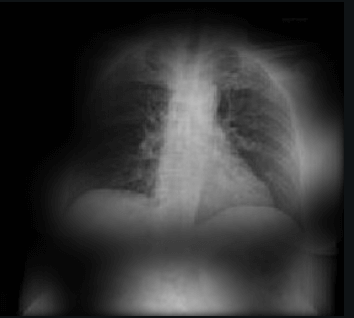 |
As noted before, researchers have recognized these leakage issues, because the Pediatric pneumonia x-rays are clearly different from COVID-19 x-rays. Leakage due to labels and cropping has been a long standing concern when working with medical images. A classic case was the presence of a ruler in some skin images for a classifier. Rulers are only included when a dermatologist believes there is a cause for concern. This led the image classifier to focus on the presence of a ruler, rather than studying the skin lesion.
Other common forms of leakage are invisible to the human eye like pixel distributions, brightness/contrast, alignments due to zoom/rotation etc. Using disparate datasets is always problematic because there are so many opportunities for introducing leakage through use case irrelevant artifacts that are very distinct to each class. Despite these concerns other data scientists have suggested using these images for helping to detect COVID-19, some examples include pyimagesearch and other bloggers.
RSNA Dataset
The issues with the Pediatric datasets, led us to instead try creating a new dataset with the RSNA images. This dataset is collated by the NIH and annotated by RSNA. This dataset contains x-rays of around ~26,000 patients. This dataset is available in DICOM format which is industry standard for medical image transfer. RSNA also includes adults.
The RSNA dataset is built from the stage 2 images available in the finished Kaggle challenge. This dataset contains 20672 Healthy and 6012 Pneumonia x-rays. We selected 20672 Healthy x-rays as Non-COVID-19 class and the 73 crowdsourced COVID-19 x-rays as the positive class.
We again built a classifier using a ResNet50 Featurizer on this dataset. Looking at the image embeddings, you can see it’s not as clear what the differences are between the COVID and non-COVID images.
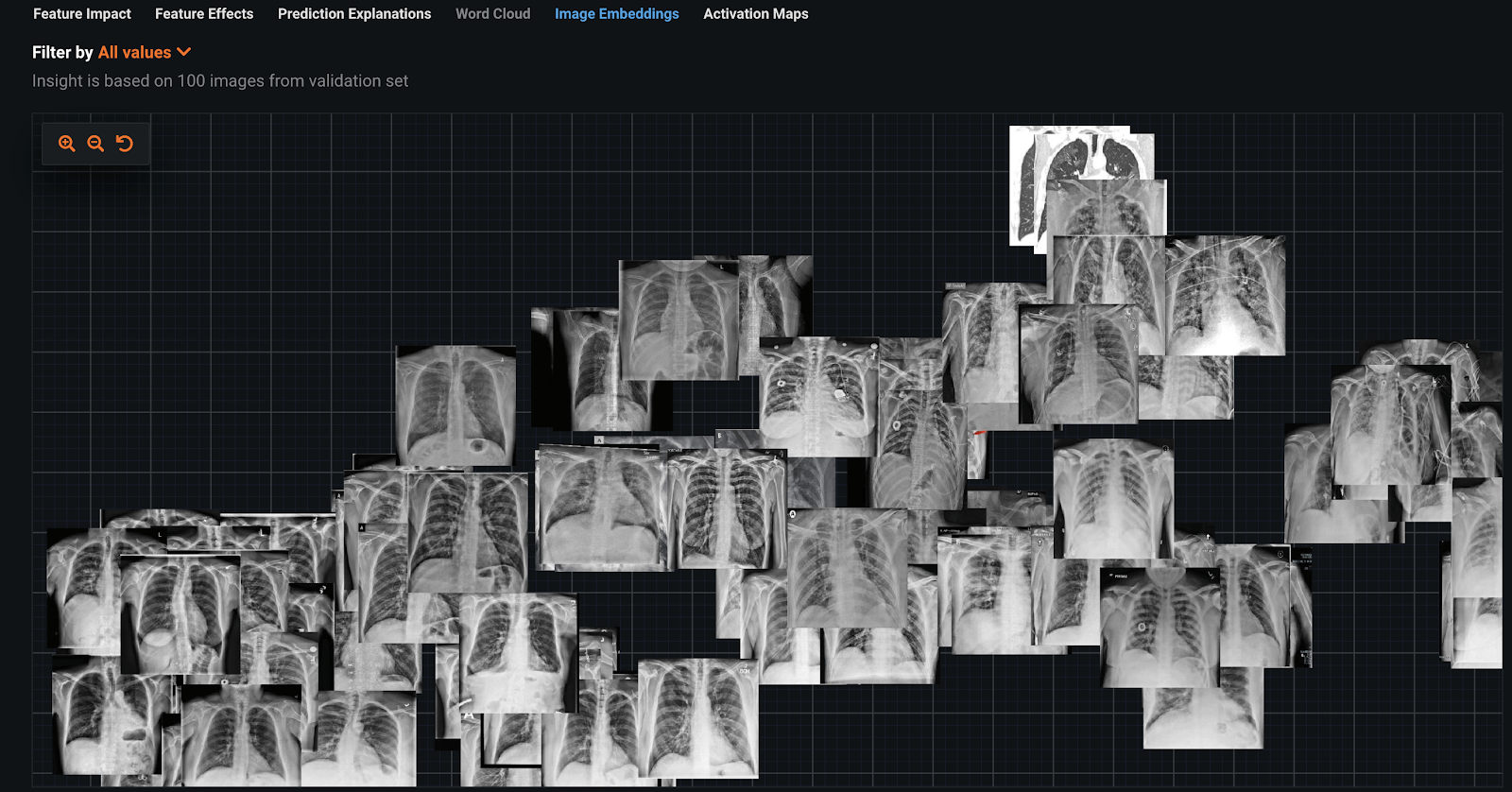
Visual AI Activation Maps highlight the areas of interest for each prediction, and the bright areas clearly show there is leakage in the dataset. Some of these issues are similar to the COVIDx dataset. Below are some examples;
Here, we can see from the bright spots in the Activation Maps that the classifier is classifying x-rays with right shoulders as COVID-19 negative. This is an example of leakage, because the classifier is focusing on how a majority of the x-rays in the RSNA dataset aren’t cropped to remove shoulders.
Here the classifier is again classifying x-rays with “L” markers by machine as COVID-19 negative. This is the same label leakage we saw earlier.
The results here show that the current datasets are not useful for detecting COVID-19 using computer vision. The RSNA and COVIDx vary from deeply flawed by using the pediatric dataset to more subtle issues of labels and cropping with the RSNA dataset with a small sample of COVID-19 positive images. A critical part of the analysis is using interpretability tools. The key to ensuring the classifiers are not falling prey to leakage was using interpretability tools like image embeddings and activation maps. While urgency is important, this experience shows that combining heterogeneous datasets for COVID-19 analysis is challenging, and that state-of-the-art interpretability techniques are mandatory to identify any issue.
Learn how DataRobot is offering services pro bono to help in the COVID-19 response effort. Also, read our blog post introducing DataRobot Visual AI for Automated Machine Learning.
About the author
Abdul Khader Jilani
Lead Execution Data Scientist
Abdul Khader Jilani is a Lead Execution Data Scientist at DataRobot. Abdul develops end-to-end enterprise AI solutions with DataRobot Enterprise AI Platform for customers across industry verticals. Before DataRobot, he was a Principal Data Scientist in Microsoft and Computer Associates, Inc.